How to Check Data Frame and Report Sizes within a Project
When you create many data frames and reports within a single project, questions may arise such as “Which data frame is occupying the most capacity?” or “Which data frame does that dashboard reference, and what is its size?”
In this note, we will explain how to use R scripts to retrieve this information as a data frame.
Two Scripts
The two R scripts used in this note are as follows:
- get_dataframe_sizes
- get_report_dataframe
get_dataframe_sizes() is a function that calculates the
disk usage of all data frames in the project and returns a data frame
sorted by total size in descending order.
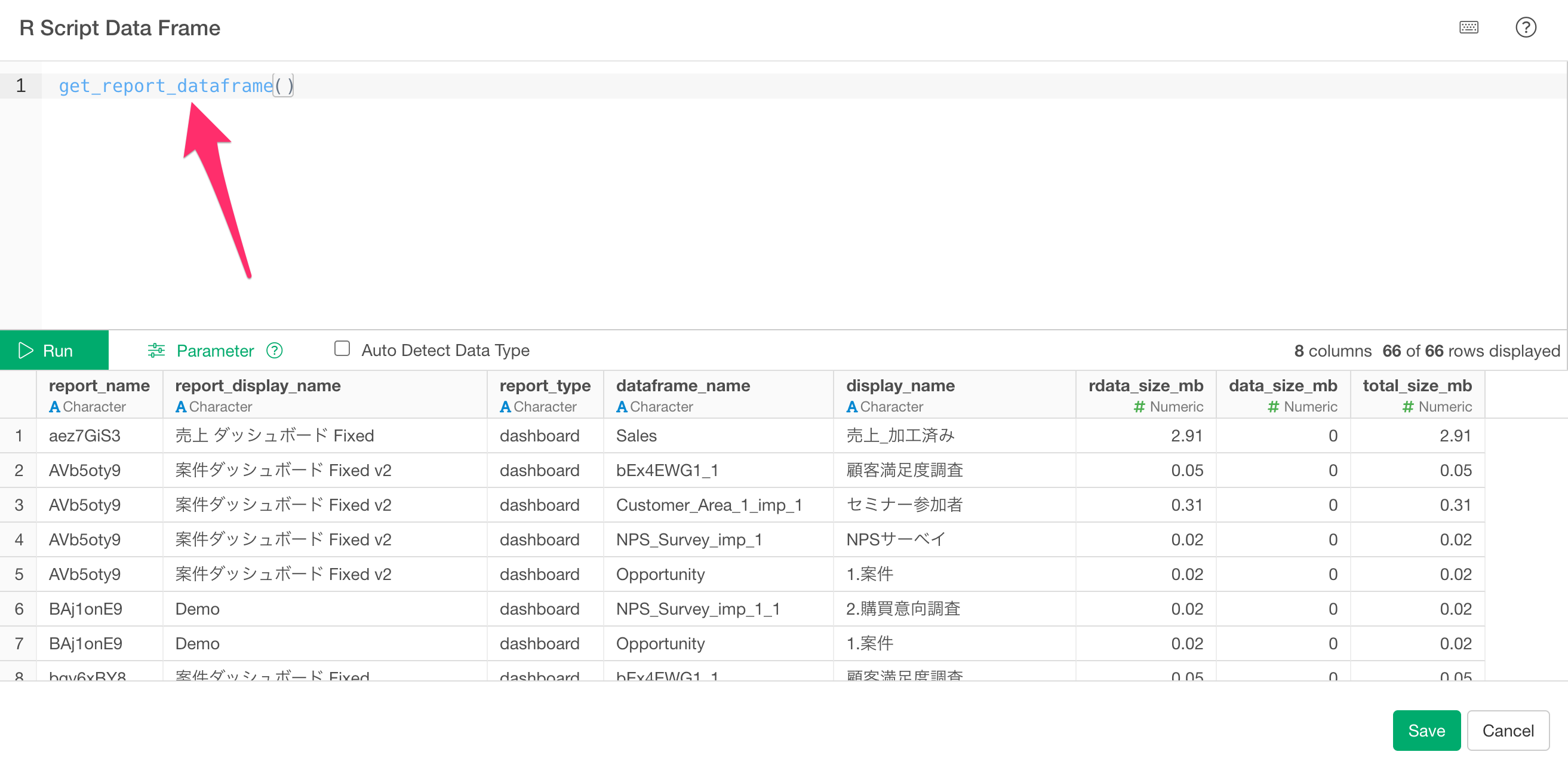
get_report_dataframe() is a function that returns a list
of which data frames are referenced by the dashboards and notes within
the project. You can check the size information of the data frames
associated with each report type.
Both functions can be called without any arguments.
Registering the R Scripts
First, register the R script file code as a script in your project.
Select “Scripts” from the “Scripts” menu at the top of the project screen.
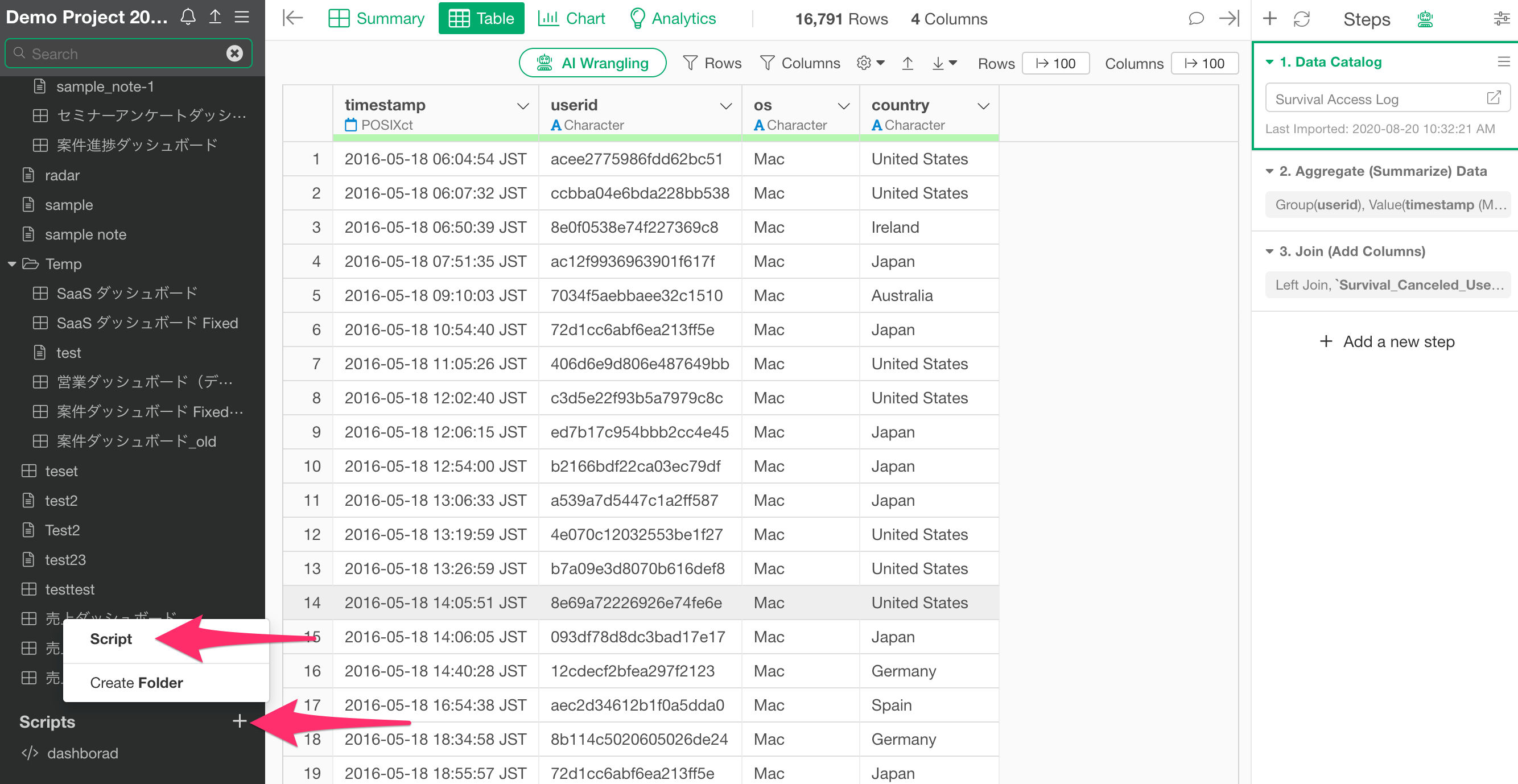
Once the script editor opens, set an arbitrary script name (e.g.,
size_check).
Next, paste the following content into the editor:
library(jsonlite)
library(dplyr)
.load_json <- function(path) {
tryCatch(
jsonlite::fromJSON(path, simplifyVector = FALSE),
error = function(e) NULL
)
}
.load_dataframes_meta <- function(project_dir) {
df_dir <- file.path(project_dir, "dataframes")
files <- list.files(df_dir, pattern = "\\.json$", full.names = TRUE)
meta <- list()
for (f in files) {
data <- .load_json(f)
if (is.null(data) || is.null(data[["name"]])) next
belongs_to <- data[["belongsTo"]][["id"]]
display_name <- data[["displayName"]]
if (is.null(display_name) || !nzchar(display_name)) display_name <- data[["name"]]
meta[[data[["name"]]]] <- list(
display_name = display_name,
belongs_to = belongs_to
)
}
meta
}
.build_step_to_root <- function(meta) {
cache <- new.env(hash = TRUE, parent = emptyenv())
resolve_root <- function(name, visited = character(0)) {
if (exists(name, envir = cache, inherits = FALSE)) {
return(get(name, envir = cache, inherits = FALSE))
}
if (name %in% visited) return(name)
df <- meta[[name]]
if (is.null(df) || is.null(df[["belongs_to"]])) {
assign(name, name, envir = cache)
return(name)
}
visited <- c(visited, name)
root <- resolve_root(df[["belongs_to"]], visited)
assign(name, root, envir = cache)
root
}
for (name in names(meta)) resolve_root(name)
as.list(cache)
}
.calculate_sizes <- function(project_dir, step_to_root, meta) {
roots <- names(meta)[sapply(names(meta), function(n) is.null(meta[[n]][["belongs_to"]]))]
sizes <- data.frame(
dataframe_name = roots,
display_name = sapply(roots, function(n) meta[[n]][["display_name"]]),
rdata_bytes = 0L,
data_bytes = 0L,
stringsAsFactors = FALSE
)
rdata_dir <- file.path(project_dir, "rdata")
for (f in list.files(rdata_dir, pattern = "\\.parquet$", full.names = TRUE)) {
stem <- tools::file_path_sans_ext(basename(f))
root <- step_to_root[[stem]]
if (!is.null(root) && root %in% roots) {
idx <- which(sizes$dataframe_name == root)
sizes$rdata_bytes[idx] <- sizes$rdata_bytes[idx] + file.info(f)$size
}
}
data_dir <- file.path(project_dir, "data")
if (dir.exists(data_dir)) {
all_files <- list.files(data_dir, full.names = TRUE)
all_files <- all_files[!file.info(all_files)$isdir]
for (f in all_files) {
stem <- tools::file_path_sans_ext(basename(f))
root <- step_to_root[[stem]]
if (!is.null(root) && root %in% roots) {
idx <- which(sizes$dataframe_name == root)
sizes$data_bytes[idx] <- sizes$data_bytes[idx] + file.info(f)$size
}
}
}
sizes$rdata_size_mb <- round(sizes$rdata_bytes / 1024^2, 2)
sizes$data_size_mb <- round(sizes$data_bytes / 1024^2, 2)
sizes$total_size_mb <- round(sizes$rdata_size_mb + sizes$data_size_mb, 2)
sizes
}
get_dataframe_sizes <- function(project_dir = dirname(getwd())) {
project_dir <- normalizePath(project_dir, mustWork = TRUE)
meta <- .load_dataframes_meta(project_dir)
step_to_root <- .build_step_to_root(meta)
sizes <- .calculate_sizes(project_dir, step_to_root, meta)
sizes |>
select(dataframe_name, display_name, rdata_size_mb, data_size_mb, total_size_mb) |>
arrange(desc(total_size_mb))
}
get_report_dataframe <- function(project_dir = dirname(getwd())) {
project_dir <- normalizePath(project_dir, mustWork = TRUE)
meta <- .load_dataframes_meta(project_dir)
step_to_root <- .build_step_to_root(meta)
sizes <- .calculate_sizes(project_dir, step_to_root, meta)
markdowns_dir <- file.path(project_dir, "markdowns")
layout_dir <- file.path(project_dir, "layout")
rows <- list()
for (md_file in sort(list.files(markdowns_dir, pattern = "\\.json$", full.names = TRUE))) {
md <- .load_json(md_file)
if (is.null(md)) next
report_name <- if (!is.null(md[["name"]])) md[["name"]] else tools::file_path_sans_ext(basename(md_file))
report_display <- md[["displayName"]]
if (is.null(report_display) || !nzchar(report_display)) report_display <- report_name
report_type <- if (isTRUE(md[["_isDashboard"]])) "dashboard" else "note"
viz_ids <- character(0)
if (!is.null(md[["dependencies"]])) {
for (dep in md[["dependencies"]]) {
if (!is.null(dep[["type"]]) && dep[["type"]] == "viz" && !is.null(dep[["id"]])) {
viz_ids <- c(viz_ids, dep[["id"]])
}
}
}
df_names <- character(0)
for (viz_id in viz_ids) {
viz_file <- file.path(layout_dir, paste0(viz_id, ".json"))
if (!file.exists(viz_file)) next
viz <- .load_json(viz_file)
if (is.null(viz) || is.null(viz[["dependencies"]])) next
for (dep in viz[["dependencies"]]) {
if (is.character(dep)) df_names <- c(df_names, dep)
}
}
df_names <- sort(unique(df_names))
for (df_name in df_names) {
size_row <- sizes[sizes$dataframe_name == df_name, ]
rows <- c(rows, list(data.frame(
report_name = report_name,
report_display_name = report_display,
report_type = report_type,
dataframe_name = df_name,
display_name = if (nrow(size_row) > 0) size_row$display_name else df_name,
rdata_size_mb = if (nrow(size_row) > 0) size_row$rdata_size_mb else 0.0,
data_size_mb = if (nrow(size_row) > 0) size_row$data_size_mb else 0.0,
total_size_mb = if (nrow(size_row) > 0) size_row$total_size_mb else 0.0,
stringsAsFactors = FALSE
)))
}
}
if (length(rows) == 0) {
return(data.frame(
report_name = character(0), report_display_name = character(0),
report_type = character(0),
dataframe_name = character(0), display_name = character(0),
rdata_size_mb = numeric(0), data_size_mb = numeric(0), total_size_mb = numeric(0)
))
}
bind_rows(rows)
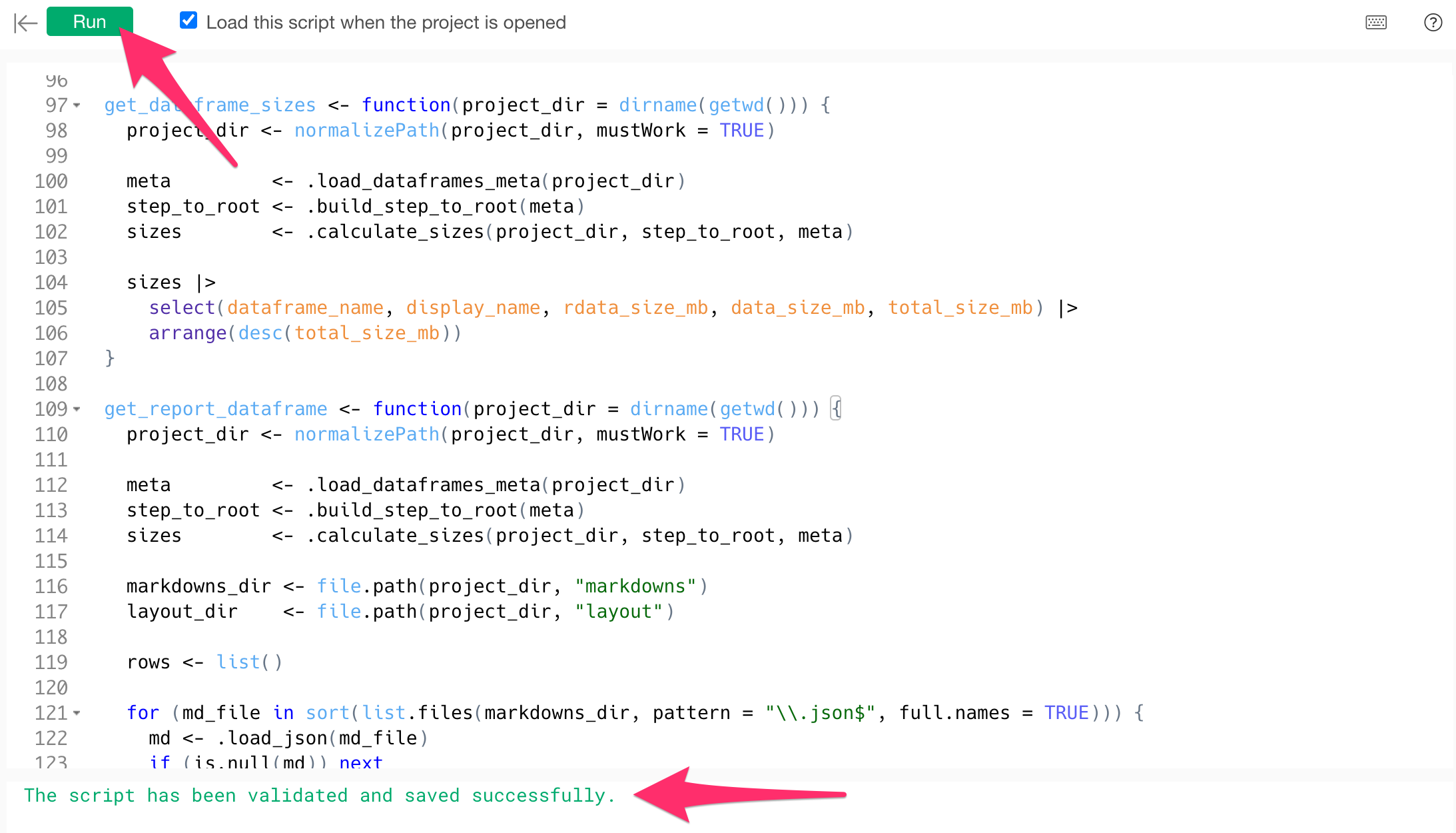
}After pasting the code, click the “Run” button. If it completes without errors, the script registration is finished.
How to Use the Functions
Checking the List of Data Frame Sizes
Click the “Add Data Frame” button and select “R Script” as the data source.
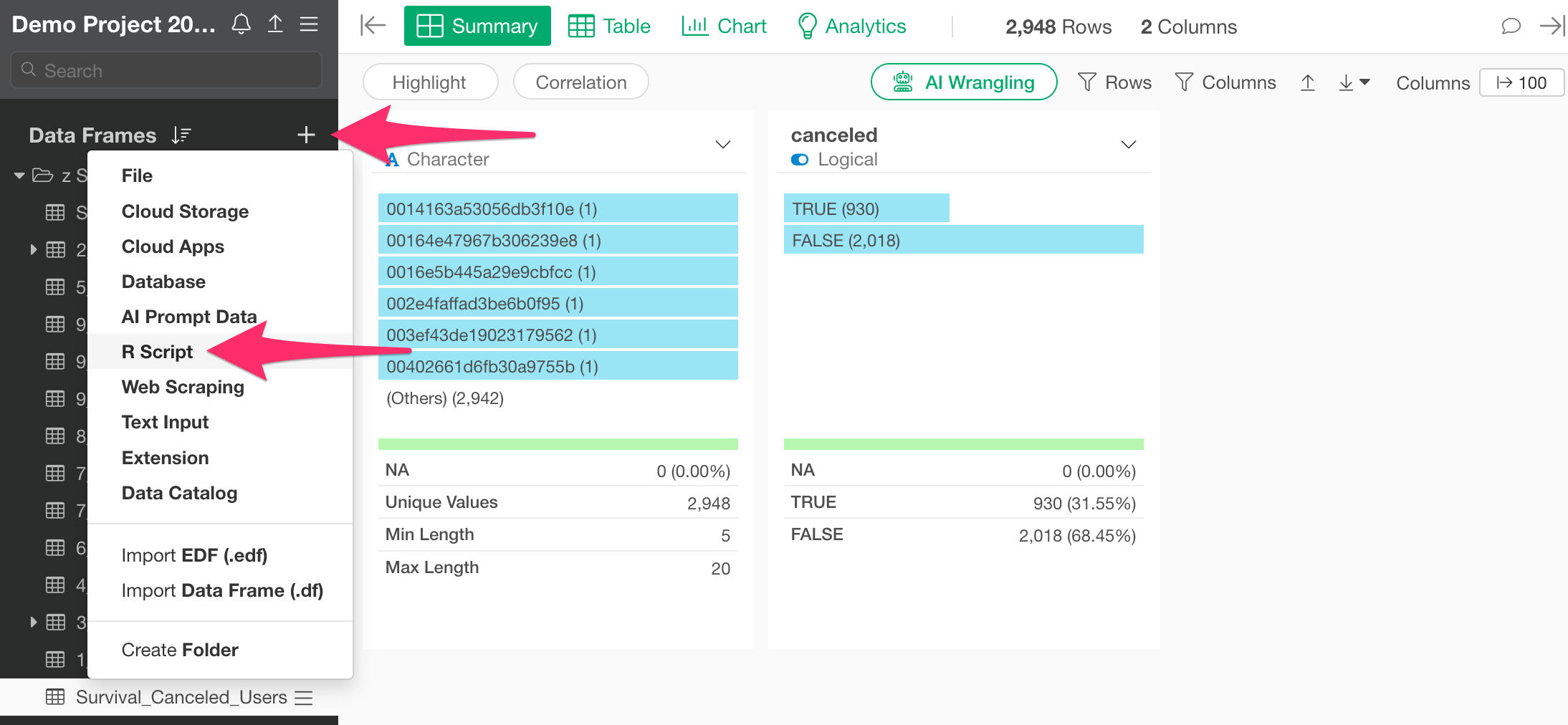
When the R script editor opens, write the following to execute the script you just registered:
get_dataframe_sizes()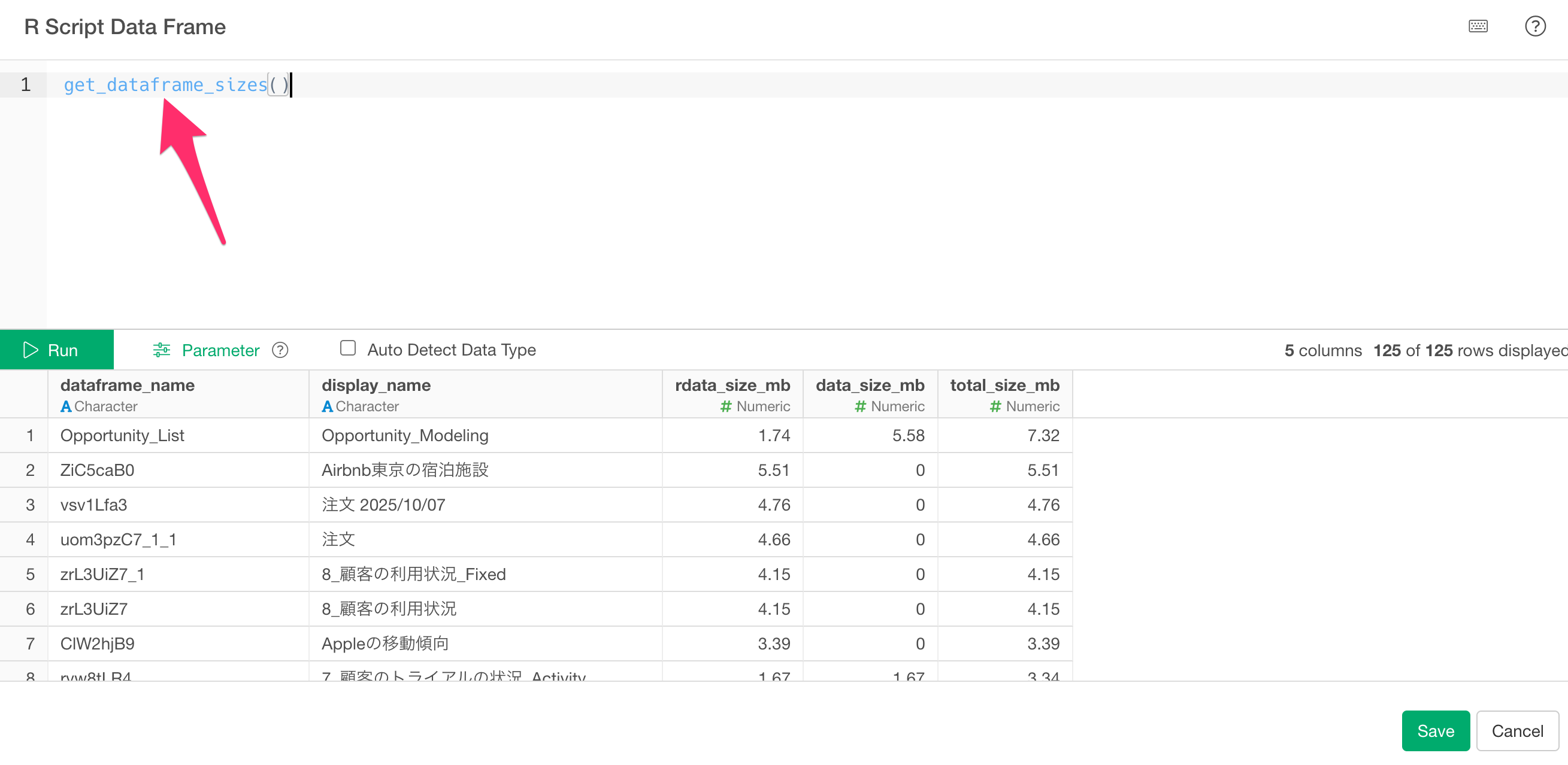
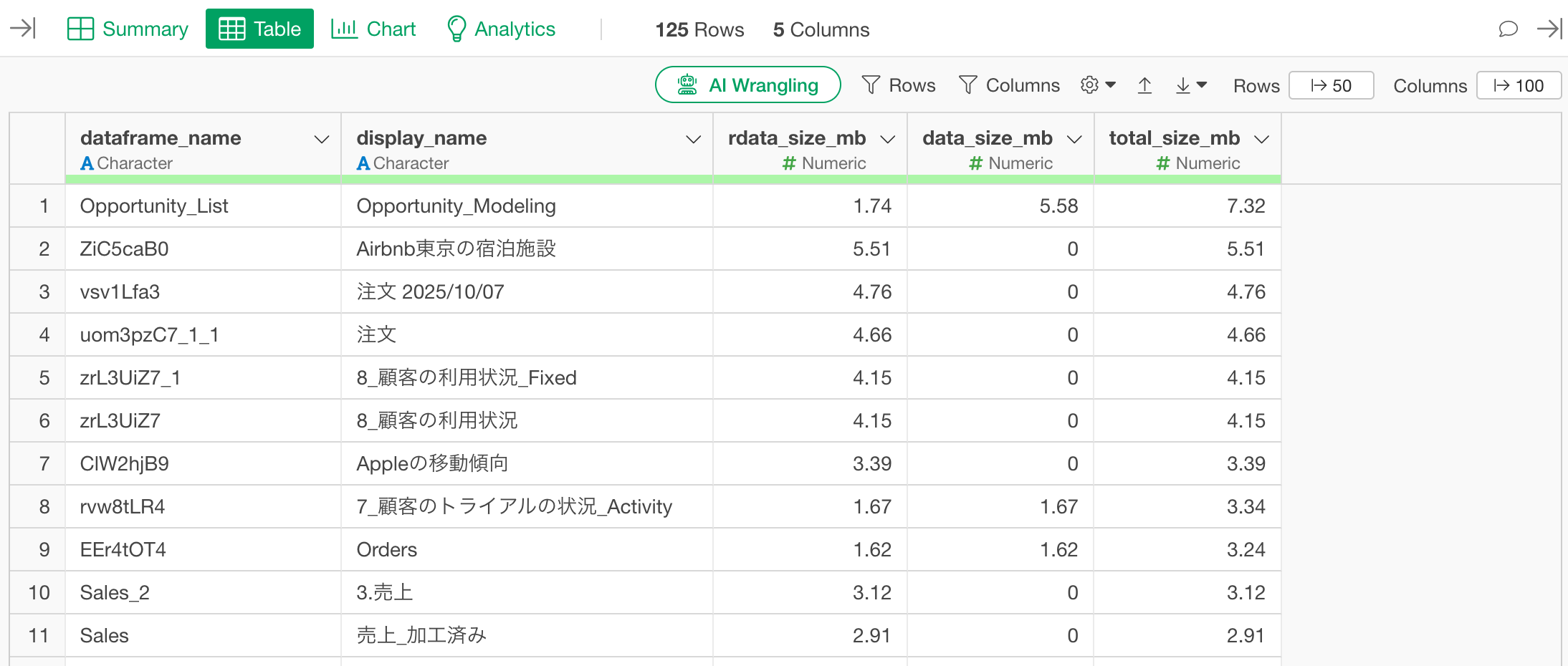
Clicking the “Run” button will return a data frame with the following columns:
- dataframe_name: Internal name of the data frame
- display_name: Name of the data frame as shown in Exploratory
- rdata_size_mb: Size of the Parquet file saved in the project’s rdata folder (MB)
- data_size_mb: Size of the original data saved in the data folder (MB)
- total_size_mb: The total size of both.
The results are sorted by total_size_mb in descending order.
By clicking the Run button, you can check the information for all data frames in the project within a data frame format.
Checking Dependencies and size of Reports and Data Frames
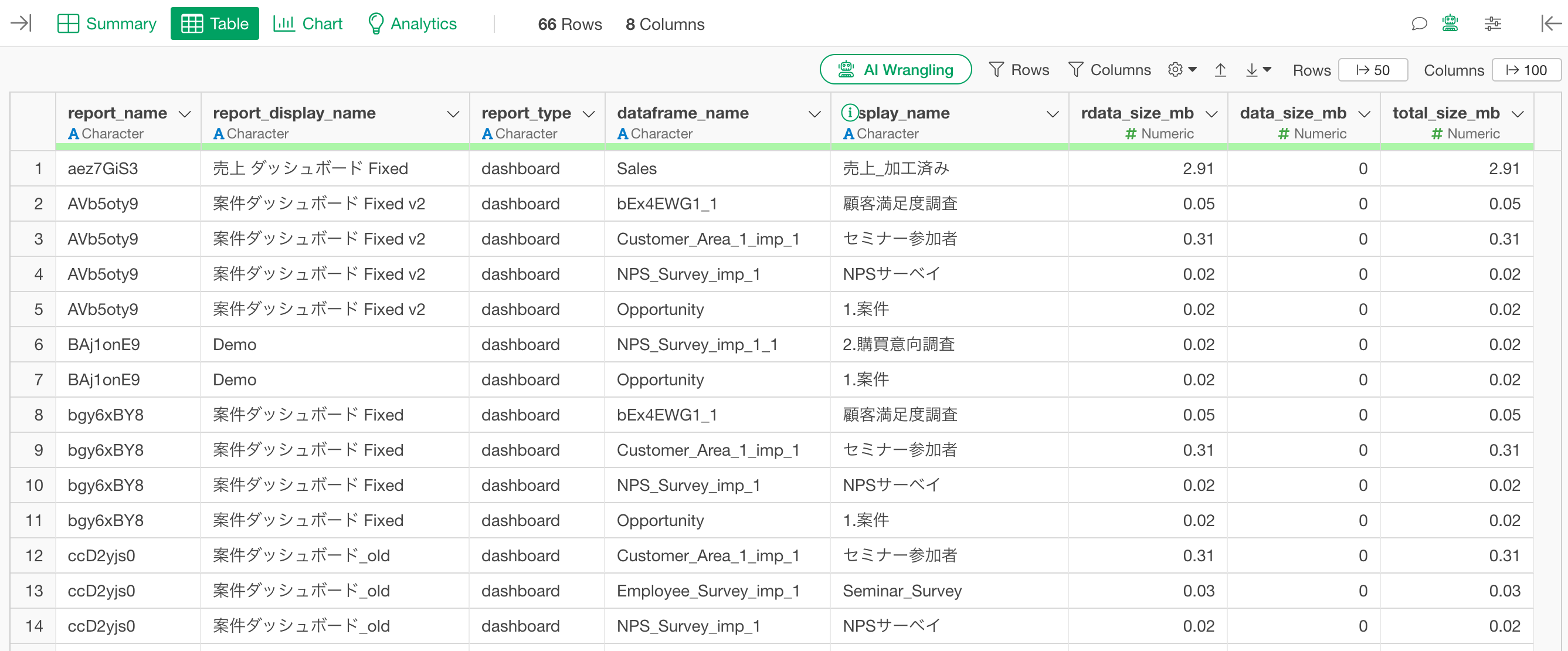
Similar to the get_dataframe_sizes function, you can use the get_report_dataframe function to see a list of which data frames each dashboard or note references, along with their sizes.
In the “R Script” data source editor, write the following:
get_report_dataframe()
Clicking the “Run” button will return a data frame with the following columns:
report_name: Internal name of the report
report_display_name: Name of the report as displayed in Exploratory
report_type: Type of report (“dashboard” or “note”)
dataframe_name: Internal name of the data frame the report depends on
display_name: Display name of that data frame
rdata_size_mb: Size of the Parquet file saved in the project’s rdata folder (MB)
data_size_mb: Size of the original data saved in the data folder (MB)
total_size_mb: The total size of both.
The results are sorted by total_size_mb in descending order.
Reference Information
- Best Practices for Improving Performance - Link